Replacing Jupyter Notebook with Org Mode
Replacing Jupyter Notebook with Org Mode
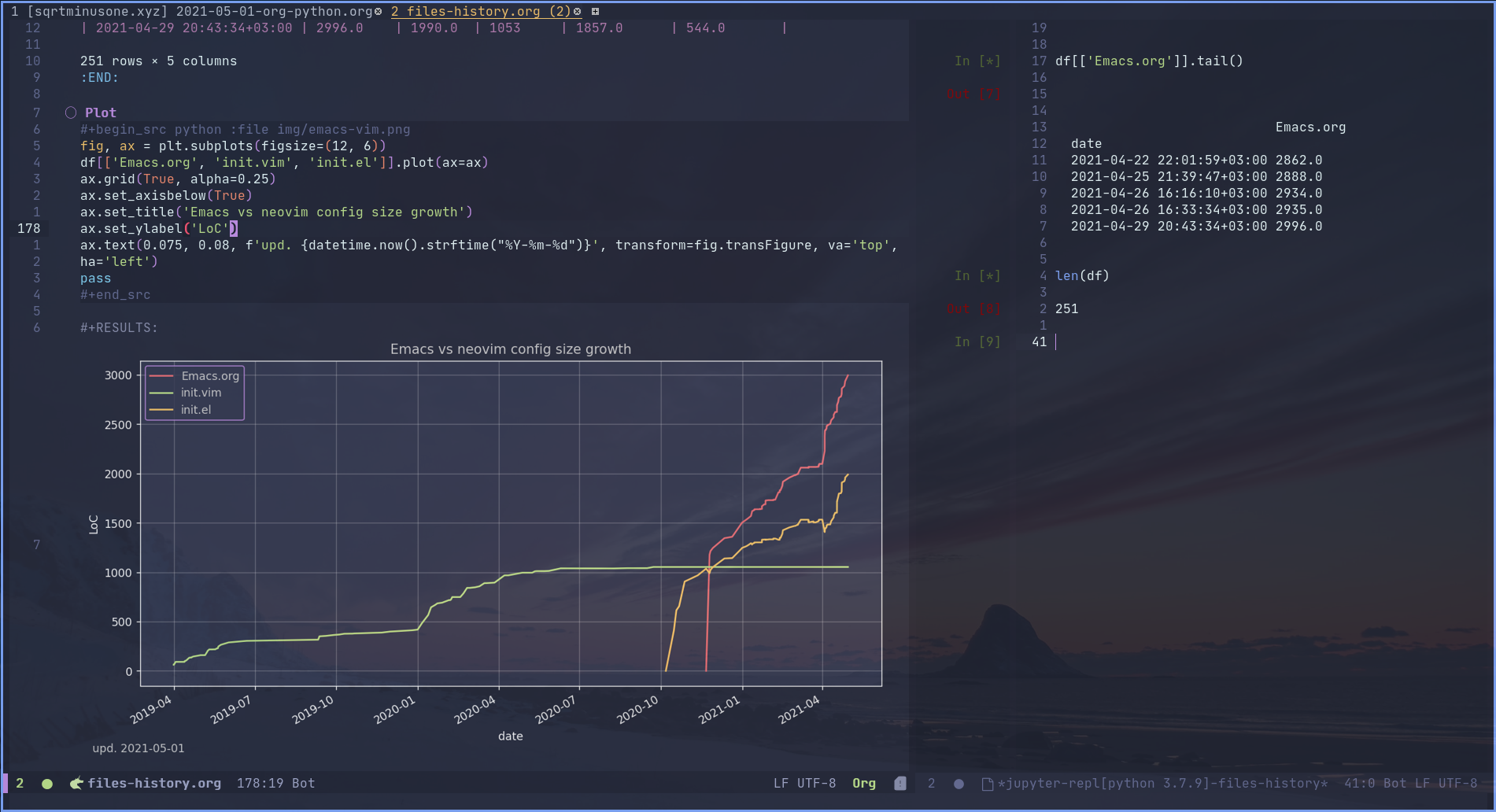
Why?
Jupyter Notebook and its successor Jupyter Lab providing an interactive development environment for many programming languages are in lots of ways great pieces of software.
But while I was using the former, and then the latter, I was also an as-full-time-as-one-can-get NeoVim user. “As one can get” is because, of course, there is no sensible way to extend the NeoVim editing experience to the Jupyter ecosystem.
A possibility for change appeared with my discovery of Emacs not so long ago. Emacs, a substantially more extensible piece of software, potentially can be used for the mentioned kind of programming. So I decided to try.
Sometime past that decision, it’s time to wrap up the results. To start with, I’ll briefly discuss the pros & cons of using Org mode rather than Jupyter Notebook/Lab. Here is my list of advantages:
- Emacs, at least for me, is way more comfortable to use than a browser
- Org mode allows using multiple programming languages in one file or multiple sessions with one programming language
- Richer & way more flexible export & tangle capacities
- More reasonable version control because org mode is just plain text, contrary to Jupyter’s JSONs
The first point deserves to be spelled out with more detail. To start with, Emacs is an objectively better text editor than Jupyter, as Emacs offers a considerable superset of Jupyter’s features concerning just writing and editing text. The farthest one can go with Jupyter Lab is to install a vim emulation plugin, which still isn’t as good as Evil mode.
The fact that Emacs can be used for different purposes also helps. For instance, I often write LaTeX documents, which are loosely based on the nearby code, e.g. using some generated charts or tables. Switching an Emacs buffer is easier than switching between Emacs and browser, not to mention that Emacs buffers usually have the same set of keybindings.
Emacs’ buffer management system, which is good enough for a window manager, is barely comparable to the tabs of Jupyter Lab. And so on.
As for why one may want to use Jupyter instead, here is my take on cons:
- Potential performance issues
- The output is not as rich as in the browser
- Collaboration with non-Emacs users is somewhat complicated
Separation of kernels, server, and client together with non-blocking JavaScript-based UI is a good argument for using Jupyter. And it certainly won’t be a problem for a browser to suddenly print a line a million characters long.
As for the richness of the output, while there are ways to work around the limitations of Emacs there, in some cases the best thing one can do is open the output in question with a browser. I’ll discuss doing that further below.
The collaboration issue can be alleviated with rich export capabilities, but if someone wants to change something in your Org file, execute it and send it back to you, they have to use Emacs.
Basic setup
The core package to this whole venture is emacs-jupyter (another notable alternative ein, using which can help with the collaboration problem).
Install it however you install packages in Emacs, here is my preferred way with use-package and straight.el:
(use-package jupyter
:straight t)
Then, we have to enable languages for org-babel. Put the following in your org mode config section:
(org-babel-do-load-languages
'org-babel-load-languages
'((emacs-lisp . t) ;; Other languages
(shell . t)
;; Python & Jupyter
(python . t)
(jupyter . t)))
Now, you should be able to use source blocks with names like jupyter-LANG, e.g. jupyter-python. To use just LANG src blocks, call the following function after org-babel-do-load-languages:
(org-babel-jupyter-override-src-block "python")
That overrides the built-in python block with jupyter-python.
If you use ob-async, you have to set jupyter-LANG blocks as ignored by this package, because emacs-jupyter has async execution of its own.
(setq ob-async-no-async-languages-alist '("python" "jupyter-python"))
Environments
So, we’ve set up a basic emacs-jupyter configuration.
The catch here is that Jupyter should be available on Emacs startup (at the time of evaluation of the emacs-jupyter package, to be precise). That means, if you are launching Emacs with something like an application launcher, global Python & Jupyter will be used.
import sys
sys.executable
/usr/bin/python3
Which is probably not what we want. To resolve that, we have to make the right Python available at the required time.
Anaconda
If you were using Jupyter Lab or Notebook before, there is a good chance you install it via Anaconda. If not, in a nutshell, it is a package & environment manager, which specializes in Python & R, but also supports a whole lot of stuff like Node.js. In my opinion, it is the easiest way to manage multiple Python installations if you don’t use some advanced package manager like Guix.
As one may expect, there is an Emacs package called conda.el to help working with conda environments in Emacs. We have to put it somewhere before the emacs-jupyter package and call conda-env-activate:
(use-package conda
:straight t
:config
(setq conda-anaconda-home (expand-file-name "~/Programs/miniconda3/"))
(setq conda-env-home-directory (expand-file-name "~/Programs/miniconda3/"))
(setq conda-env-subdirectory "envs"))
(unless (getenv "CONDA_DEFAULT_ENV")
(conda-env-activate "base"))
If you have Anaconda installed on a custom path, as I do, you’d have to add these setq lines in the :config section. Also, there is no point in activating the environment if Emacs is somehow already launched in an environment.
That’ll give us Jupyter from a base conda environment.
If you use a plain virtual environment, you can use virtualenvwrapper.el, which is similar in its design to conda.el (or, rather, the other way round).
Switching an environment
However, as you will notice rather soon, emacs-jupyter will always use the Python kernel found on startup. So if you switch to a new environment, the code will still be running in the old one, which is not too convenient.
Fortunately, to fix that we have only to force the refresh of Jupyter kernelspecs:
(defun my/jupyter-refresh-kernelspecs ()
"Refresh Jupyter kernelspecs"
(interactive)
(jupyter-available-kernelspecs t))
Calling M-x my/jupyter-refresh-kernelspecs after an environment switch will give you a new kernel. Just keep in mind that a kernelspec seems to be attached to a session, so you’d also have to change the session name to get a new kernel.
import sys
sys.executable
/home/pavel/Programs/miniconda3/bin/python
(conda-env-activate "ann")
import sys
sys.executable
/home/pavel/Programs/miniconda3/bin/python
(my/jupyter-refresh-kernelspecs)
import sys
sys.executable
/home/pavel/Programs/miniconda3/envs/ann/bin/python
Programming
To test if everything is working correctly, run M-x jupyter-run-repl, which should give you a REPL with a chosen kernel. If so, we can finally start using Python in org mode.
#+begin_src python :session hello :async yes
print('Hello, world!')
#+end_src
#+RESULTS:
: Hello, world!
#+end_src
To avoid repeating similar arguments for the src block, we can set the header-args property at the start of the file:
#+PROPERTY: header-args:python :session hello
#+PROPERTY: header-args:python+ :async yes
When a kernel is initialized, an associated REPL buffer is also created with a name like *jupyter-repl[python 3.9.2]-hello*.
One advantage of emacs-jupyter over the standard Org source execution is that kernel requests for input are queried through the minibuffer. So, you can run a code like this:
#+begin_src python
name = input('Name: ')
print(f'Hello, {name}!')
#+end_src
#+RESULTS:
: Hello, Pavel!
without any additional setup.
Code output
Images
Image output should work out of the box. Run M-x org-toggle-inline-images (C-c C-x C-v) after the execution to see the image inline.
#+begin_src python
import matplotlib.pyplot as plt
fig, ax = plt.subplots()
ax.plot([1, 2, 3, 4], [1, 4, 2, 3])
pass
#+end_src
#+RESULTS:
[[file:./.ob-jupyter/86b3c5e1bbaee95d62610e1fb9c7e755bf165190.png]]
There is some room for improvement though. First, you can add the following hook if you don’t want to press this awkward keybinding every time:
(add-hook 'org-babel-after-execute-hook 'org-redisplay-inline-images)
Second, we may override the image save path like this:
#+begin_src python :file img/hello.png
import matplotlib.pyplot as plt
fig, ax = plt.subplots()
ax.plot([1, 2, 3, 4], [1, 4, 2, 3])
pass
#+end_src
#+RESULTS:
[[file:img/hello.png]]
That can save you a savefig call if the image has to be used somewhere further.
Finally, by default, the image has a transparent background and a ridiculously small size. That can be fixed with some matplotlib settings:
import matplotlib as mpl
mpl.rcParams['figure.dpi'] = 200
mpl.rcParams['figure.facecolor'] = '1'
At the same time, we can set the image width to prevent images from becoming too large. I prefer to do it inside a emacs-lisp code block in the same org file:
(setq-local org-image-actual-width '(1024))
Basic tables
If you are evaluating something like pandas DataFrame, it will be outputted in the HTML format, wrapped in the begin_export block. To view the data in text format, you can set :display plain:
#+begin_src python :display plain
import pandas as pd
pd.DataFrame({"a": [1, 2], "b": [3, 4]})
#+end_src
#+RESULTS:
: a b
: 0 1 3
: 1 2 4
Another solution is to use something like the tabulate package:
#+begin_src python
import pandas as pd
import tabulate
df = pd.DataFrame({"a": [1, 2], "b": [3, 4]})
print(tabulate.tabulate(df, headers=df.columns, tablefmt="orgtbl"))
#+end_src
#+RESULTS:
: | | a | b |
: |----+-----+-----|
: | 0 | 1 | 3 |
: | 1 | 2 | 4 |
HTML & other rich output
Yet another solution is to use emacs-jupyter’s option :pandoc t, which invokes pandoc to convert HTML, LaTeX, and Markdown to Org. Predictably, this is slower than the options above.
#+begin_src python :pandoc t
import pandas as pd
df = pd.DataFrame({"a": [1, 2], "b": [3, 4]})
df
#+end_src
#+RESULTS:
:RESULTS:
| | a | b |
|---+---+---|
| 0 | 1 | 3 |
| 1 | 2 | 4 |
:END:
Also, every once in a while I have to view an actual, unconverted HTML in a browser, e.g. when using folium or displaCy.
To do that, I’ve written a small function, which performs xdg-open on the HTML export block under the cursor:
(setq my/org-view-html-tmp-dir "/tmp/org-html-preview/")
(use-package f
:straight t)
(defun my/org-view-html ()
(interactive)
(let ((elem (org-element-at-point))
(temp-file-path (concat my/org-view-html-tmp-dir (number-to-string (random (expt 2 32))) ".html")))
(cond
((not (eq 'export-block (car elem)))
(message "Not in an export block!"))
((not (string-equal (plist-get (car (cdr elem)) :type) "HTML"))
(message "Export block is not HTML!"))
(t (progn
(f-mkdir my/org-view-html-tmp-dir)
(f-write (plist-get (car (cdr elem)) :value) 'utf-8 temp-file-path)
(start-process "org-html-preview" nil "xdg-open" temp-file-path))))))
f.el is used by a lot of packages, including the above-mentioned conda.el, so you probably already have it installed.
Put a cursor on the begin_export html block and run M-x my/org-view-html.
There also seems to be widgets support in emacs-jupyter, but I wasn’t able to make it work.
DataFrames
Last but not least option I want to mention here is specifically about pandas’ DataFrames. There aren’t many good options to view the full dataframe inside Emacs. One way I can think of is to save the dataframe in CSV and view it with csv-mode.
However, there are standalone packages to view dataframes. One I can point out is dtale, which is a Flask + React app designed just for that purpose. It has a rather extensive list of features, including charting, basic statistical instruments, filters, etc. Here is an online demo.
The problem here is that it’s a browser app, which means it defies one of the purposes of using Org Mode in the first place. What’s more, this application is a rather huge one with lots of dependencies, and they have to be installed in the same environment as your project.
So this approach has its pros and cons as well. Example usage is as follows:
import dtale
d = dtale.show(df)
d.open_browser() # Or get an URL from d._url
Another notable alternative is PandasGUI, which, as one can guess, is a GUI (PyQt5) application, although it uses QtWebEngine inside.
Remote kernels
There are yet some problems in the current configuration.
- Input/output handling is far from perfect. For instance, (at least in my configuration) Emacs tends to get slow for log-like outputs, e.g. Keras with
verbose=2. It may even hang if the output is one long line. ipdbbehaves awkwardly if called from ansrcblock, although it at least will let you typequit.- Whenever you close Emacs, kernels are stopped, so you’d have to execute the code again on the next start.
Using a “remote” kernel
For the reasons above I sometimes prefer to use a standalone kernel. To start a Jupyter kernel, run the following command in the environment and path you need:
jupyter kernel --kernel=python
After the kernel is launched, write the path to the connection file into the :session header and press C-c C-c to refresh the setup:
#+PROPERTY: header-args:python :session /home/pavel/.local/share/jupyter/runtime/kernel-e770599c-2c98-429b-b9ec-4d1ddf5fc16c.json
Now python source blocks should be executed in the kernel.
To open a REPL, run M-x jupyter-connect-repl and select the given JSON. Or launch a standalone REPL like this:
jupyter qtconsole --existing kernel-e770599c-2c98-429b-b9ec-4d1ddf5fc16c.json
Executing a piece of code in the REPL allows proper debugging, for instance with %pdb magic. Also, Jupyter QtConsole generally handles large outputs better and even allows certain kinds of rich output in the REPL.
Some automation
Now, I wouldn’t use Emacs if it wasn’t possible to automate at least some of the listed steps. So here are the functions I’ve written for that.
First, we need to get open ports on the system:
(defun my/get-open-ports ()
(mapcar
#'string-to-number
(split-string (shell-command-to-string "ss -tulpnH | awk '{print $5}' | sed -e 's/.*://'") "\n")))
Then, list the available kernel JSONs:
(setq my/jupyter-runtime-folder (expand-file-name "~/.local/share/jupyter/runtime"))
(defun my/list-jupyter-kernel-files ()
(mapcar
(lambda (file) (cons (car file) (cdr (assq 'shell_port (json-read-file (car file))))))
(sort
(directory-files-and-attributes my/jupyter-runtime-folder t ".*kernel.*json$")
(lambda (x y) (not (time-less-p (nth 6 x) (nth 6 y)))))))
And query the user for a running kernel:
(defun my/select-jupyter-kernel ()
(let ((ports (my/get-open-ports))
(files (my/list-jupyter-kernel-files)))
(completing-read
"Jupyter kernels: "
(seq-filter
(lambda (file)
(member (cdr file) ports))
files))))
After which we can use the my/select-jupyter-kernel function however we want:
(defun my/insert-jupyter-kernel ()
"Insert a path to an active Jupyter kernel into the buffer"
(interactive)
(insert (my/select-jupyter-kernel)))
(defun my/jupyter-connect-repl ()
"Open emacs-jupyter REPL, connected to a Jupyter kernel"
(interactive)
(jupyter-connect-repl (my/select-jupyter-kernel) nil nil nil t))
(defun my/jupyter-qtconsole ()
"Open Jupyter QtConsole, connected to a Jupyter kernel"
(interactive)
(start-process "jupyter-qtconsole" nil "setsid" "jupyter" "qtconsole" "--existing"
(file-name-nondirectory (my/select-jupyter-kernel))))
The first function, which simply inserts the path to the kernel, is meant to be used on the :session header. One can go even further and locate the header automatically, but that’s an idea for next time.
The second one opens a REPL provided by emacs-jupyter. The t argument is necessary to pop up the REPL immediately.
The last one launches Jupyter QtConsole. setsid is required to run the program in a new session, so it won’t close together with Emacs.
Cleaning up
I’ve also noticed that there are JSON files left in the runtime folder whenever the kernel isn’t stopped correctly. So here is a cleanup function.
(defun my/jupyter-cleanup-kernels ()
(interactive)
(let* ((ports (my/get-open-ports))
(files (my/list-jupyter-kernel-files))
(to-delete (seq-filter
(lambda (file)
(not (member (cdr file) ports)))
files)))
(when (and (length> to-delete 0)
(y-or-n-p (format "Delete %d files?" (length to-delete))))
(dolist (file to-delete)
(delete-file (car file)))))
Export
An uncountable number of articles have been written already on the subject of Org Mode export, so I will just cover my particular setup.
HTML
Export to a standalone HTML is an easy way to share the code with someone who doesn’t use Emacs, just remember that HTML may not be the only file you’d have to share if you have images in the document. Although you may use something like htmlark later to get a proper self-contained HTML.
To do the export, run M-x org-html-export-to-html. It should work out of the box, however, we can improve the output a bit.
First, we can add a custom CSS to the file. I like this one:
#+HTML_HEAD: <link rel="stylesheet" type="text/css" href="https://gongzhitaao.org/orgcss/org.css"/>
To get a syntax highlighting, we need the htmlize package:
(use-package htmlize
:straight t
:after ox
:config
(setq org-html-htmlize-output-type 'css))
If you use the rainbow-delimeters package, as I do, default colors for delimiters may not look good with the light theme. To fix such issues, put an HTML snippet like this in a begin_export html block or a CSS file:
<style type="text/css">
.org-rainbow-delimiters-depth-1, .org-rainbow-delimiters-depth-2, .org-rainbow-delimiters-depth-3, .org-rainbow-delimiters-depth-4 {
color: black
}
</style>
LaTeX -> pdf
Even though I use LaTeX quite extensively, I don’t like to add another layer of complexity here and 98% of the time write plain .tex files. LaTeX by itself provides many good options whenever you need to write a document together with some data or source code, contrary to “traditional” text processors.
Nevertheless, I want to get at least a tolerable pdf from Org, so here is a piece of my config with some inline comments.
(defun my/setup-org-latex ()
(setq org-latex-compiler "xelatex") ;; Probably not necessary
(setq org-latex-pdf-process '("latexmk -outdir=%o %f")) ;; Use latexmk
(setq org-latex-listings 'minted) ;; Use minted to highlight source code
(setq org-latex-minted-options ;; Some minted options I like
'(("breaklines" "true")
("tabsize" "4")
("autogobble")
("linenos")
("numbersep" "0.5cm")
("xleftmargin" "1cm")
("frame" "single")))
;; Use extarticle without the default packages
(add-to-list 'org-latex-classes
'("org-plain-extarticle"
"\\documentclass{extarticle}
[NO-DEFAULT-PACKAGES]
[PACKAGES]
[EXTRA]"
("\\section{%s}" . "\\section*{%s}")
("\\subsection{%s}" . "\\subsection*{%s}")
("\\subsubsection{%s}" . "\\subsubsection*{%s}")
("\\paragraph{%s}" . "\\paragraph*{%s}")
("\\subparagraph{%s}" . "\\subparagraph*{%s}"))))
;; Make sure to eval the function when org-latex-classes list already exists
(with-eval-after-load 'ox-latex
(my/setup-org-latex))
In the document itself, add the following headers:
#+LATEX_CLASS: org-plain-extarticle
#+LATEX_CLASS_OPTIONS: [a4paper, 14pt]
14pt size is required by certain state standards of ours for some reason.
After which you can put whatever you want in the preamble with LATEX_HEADER. My workflow with LaTeX is to write a bunch of .sty files beforehand and import the necessary ones in the preamble. Here is the repo with these files, although quite predictably, it’s a mess. At any rate, I have to write something like the following in the target Org file:
#+LATEX_HEADER: \usepackage{styles/generalPreamble}
#+LATEX_HEADER: \usepackage{styles/reportFormat}
#+LATEX_HEADER: \usepackage{styles/mintedSourceCode}
#+LATEX_HEADER: \usepackage{styles/russianLocale}
M-x org-latex-export-to-latex should export the document to the .tex file. As an alternative, run M-x org-export-dispatch (by default should be on C-c C-e) an pick the required option there.
ipynb
One last export backend I want to mention is ox-ipynb, which allows exporting Org documents to Jupyter notebooks. Sometimes it works, sometimes it doesn’t.
Also, the package isn’t on MELPA, so you have to install it from the repo directly.
(use-package ox-ipynb
:straight (:host github :repo "jkitchin/ox-ipynb")
:after ox)
To (try to) do export, run M-x ox-ipynb-export-org-file-ipynb-file.